Nuclear Factor Y (NF-Y)
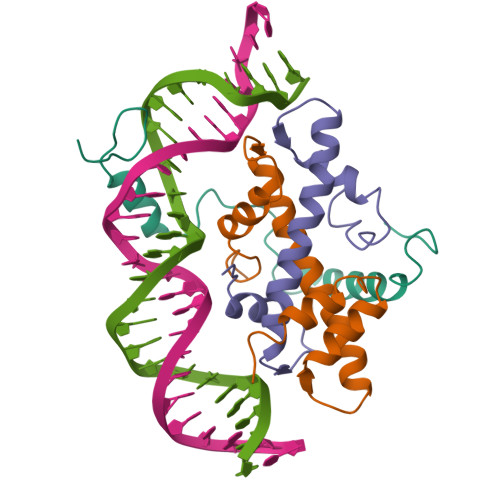
The sequence-specific transcription factor NF-Y binds the CCAAT box, one of the most frequently occurring regulatory elements in eukaryotic promoters. NF-Y functions as a heterotrimer composed of the NF-YA subunit and the NF-YB/NF-YC dimer, the latter two containing a conserved histone-fold domain (HFD).
In 2013, we reported the first crystal structure of the NF-Y/DNA complex, consisting of the NF-Y trimer bound to a 25-bp DNA fragment containing a CCAAT box (PDB code: 4AWL). This structure revealed the molecular basis of a sequence-specific, histone-like mode of DNA recognition by NF-Y (Nardini et al., 2013). NF-YA mediates both trimer assembly and DNA binding by inserting an α-helix deeply into the DNA minor groove, thereby establishing sequence-specific contacts with the CCAAT motif. Structural analysis combined with mutational data demonstrated that ubiquitination of NF-YB at Lys138 precedes and functionally mirrors monoubiquitination of histone H2B at Lys120, a key modification in transcriptional activation. These findings identify NF-Y as a pioneer transcription factor with nucleosome-like properties that promotes permissive chromatin states at CCAAT-containing promoters. They further suggest that other HFD-containing proteins may operate through similar mechanisms (Gnesutta et al., 2013; Nardone et al., 2017).
NF-Y is known to cooperate with neighboring TFs to regulate gene expression. Genome-wide analyses detected a precise stereo-alignment -10/12 bp- of CCAAT with E-box elements and corresponding colocalization of NF-Y with basic-Helix-Loop-Helix (bHLH) TFs. We dissected NF-Y interactions with USF1 and MAX. USF1, but not MAX, cooperates in DNA binding with NF-Y. NF-Y and USF1 synergize to activate target promoters. Reconstruction of complexes by structural means shows independent DNA binding of MAX, whereas USF1 has extended contacts with NF-Y, involving the USR, a USF-specific amino acid sequence stretch required for trans-activation. The USR is an intrinsically disordered domain and adopts different conformations based on E-box-CCAAT distances. Deletion of the USR abolishes cooperative DNA binding with NF-Y. Our data indicate that the functionality of certain unstructured domains involves adapting to small variation in stereo-alignments of the multimeric TFs sites (Bernardini et al., 2021).
In plants, NF-Y has undergone extensive genetic diversification, with multiple genes encoding each subunit. This expansion enables the formation of numerous NF-Y heterotrimeric complexes, providing a flexible and highly combinatorial system capable of regulating distinct developmental and physiological pathways. To date, we have determined the structure of the Arabidopsis thaliana NF-YA6/NF-YB2/NF-YC3 complex bound to its target FT CCAAT DNA element (PDB code: 6R2V), as well as several NF-YB/NF-YC histone-like dimers, including NF-YB6(L1L)/NF-YC3 (PDB code: 5G49), NF-YB2/NF-YC3 (PDB codes: 6R0M and 6R0N), and the rice complex Ghd8/OsNF-YC7 (PDB code: 6R0L) (Chaves-Sanjuan et al., 2020; Gnesutta et al., 2017).
NF-Y plays also a critical role in cancer biology. Tumor cells frequently undergo metabolic rewiring through the overexpression of genes involved in biosynthetic and metabolic pathways, many of which are directly regulated by NF-Y. Consistently, NF-YA is overexpressed in multiple tumor types. These observations highlight NF-Y as a potential therapeutic target, and efforts to modulate its activity are an active area of investigation. Within this context, we have identified suramin as a promising small-molecule inhibitor of NF-Y function (Nardone et al., 2020).
Publications
Nardini M, Gnesutta N, Donati G, Gatta R, Forni C, Fossati A, Vonrhein C, Moras D, Romier C, Bolognesi M, Mantovani R. Sequence-specific transcription factor NF-Y displays histone-like DNA binding and H2B-like ubiquitination.
Cell. 2013 Jan 17;152(1-2):132-43. doi: 10.1016/j.cell.2012.11.047. [PubMed: 23332751]
Gnesutta N, Nardini M, Mantovani R. The H2A/H2B-like histone-fold domain proteins at the crossroad between chromatin and different DNA metabolisms.
Transcription. 2013 May-Jun;4(3):114-9. [PubMed: 23756340]
Gnesutta N, Saad D, Chaves-Sanjuan A, Mantovani R, Nardini M. Crystal Structure of the Arabidopsis thaliana L1L/NF-YC3 Histone-fold Dimer Reveals Specificities of the LEC1 Family of NF-Y Subunits in Plants.
Mol Plant. 2017 Apr 3;10(4), 645-648. doi: 10.1016/j.molp.2016.11.006 [PubMed: 27871811]
Nardone V, Chaves-Sanjuan A, Nardini M. Structural determinants for NF-Y/DNA interaction at the CCAAT box.
Biochim Biophys Acta. 2017 May;1860(5):571-580. doi: 10.1016/j.bbagrm.2016.09.006 [PubMed: 27677949]
Bernardini A, Lorenzo M, Nardini M, Mantovani R, Gnesutta N. The phosphorylatable Ser320 of NF-YA is involved in DNA binding of the NF-Y trimer.
FASEB J. 2019 Apr;33(4):4790-4801. doi: 10.1096/fj.201801989R. [PubMed: 30589568]
Nardone V, Chaves-Sanjuan A, Lapi M, Airoldi C, Saponaro A, Pasqualato S, Dolfini D, Camilloni C, Bernardini A, Gnesutta N, Mantovani R, Nardini M. Structural basis of inhibition of the pioneer transcription factor NF-Y by suramin.
Cells. 2020 Oct 29;9(11):2370. doi: 10.3390/cells9112370. [PubMed: 33138093]
Chaves-Sanjuan A, Gnesutta N, Gobbini A, Martignago D, Bernardini A, Fornara F, Mantovani R, Nardini M. Structural determinants for NF-Y subunit organization and NF-Y/DNA association in plants.
Plant J. 2021 Jan;105(1):49-61. doi: 10.1111/tpj.15038. [PubMed: 33098724]
Bernardini A, Lorenzo M, Chaves-Sanjuan A, Swuec P, Pigni M, Saad D, Konarev PV, Graewert MA, Valentini E, Svergun DI, Nardini M, Mantovani R, Gnesutta N. The USR domain of USF1 mediates NF-Y interactions and cooperative DNA binding.
Int J Biol Macromol. 2021 Dec 15;193(Pt A):401-413. doi: 10.1016/j.ijbiomac.2021.10.056. [PubMed: 34673109]
Lab Team
Haidi Shahi, Antonio Chaves-Sanjuan, Louise Gourlay, Marco Nardini
Main collaborators
Prof. R. Mantovani and Prof. N. Gnesutta, Dept. of Biosciences, University of Milano (Italy)
